Acute Kidney Injury Predictor
Will your patient develop AKI?
Acute Kidney Injury (AKI) occurs frequently in critically ill patients. AKI is associated with increased morbidity and mortality, and a high financial cost. The Kidney Disease/Improving Global Outcomes (KDIGO) working group has defined and classified AKI in three stages of increasing severity, according to serum creatinine levels and urine output. Although no effective treatment currently exists that could attenuate the course of AKI, early prediction of AKI to detect the patients at risk could be a first step in the discovery and assessment of new therapies. For that purpose, we propose an AKI predictor for patients admitted to an intensive care unit (ICU). Using routinely available patient characteristics, the AKI predictor is able to predict if an adult patient will develop any stage of AKI (as defined by the KDIGO serum creatinine criteria) during the first week of ICU stay.
Project description
The AKI predictor is based on extensive statistical retrospective analysis of data from a large number of adult ICU patients. The database of the large prospective multicenter randomized EPaNIC clinical trial1 was used to build and validate the predictive models. In this study, 4640 patients were randomized to receive either early or late parenteral nutrition. The models were constructed using Random Forests, a data mining technique. They have good discrimination and are well calibrated in this large cohort of patients.
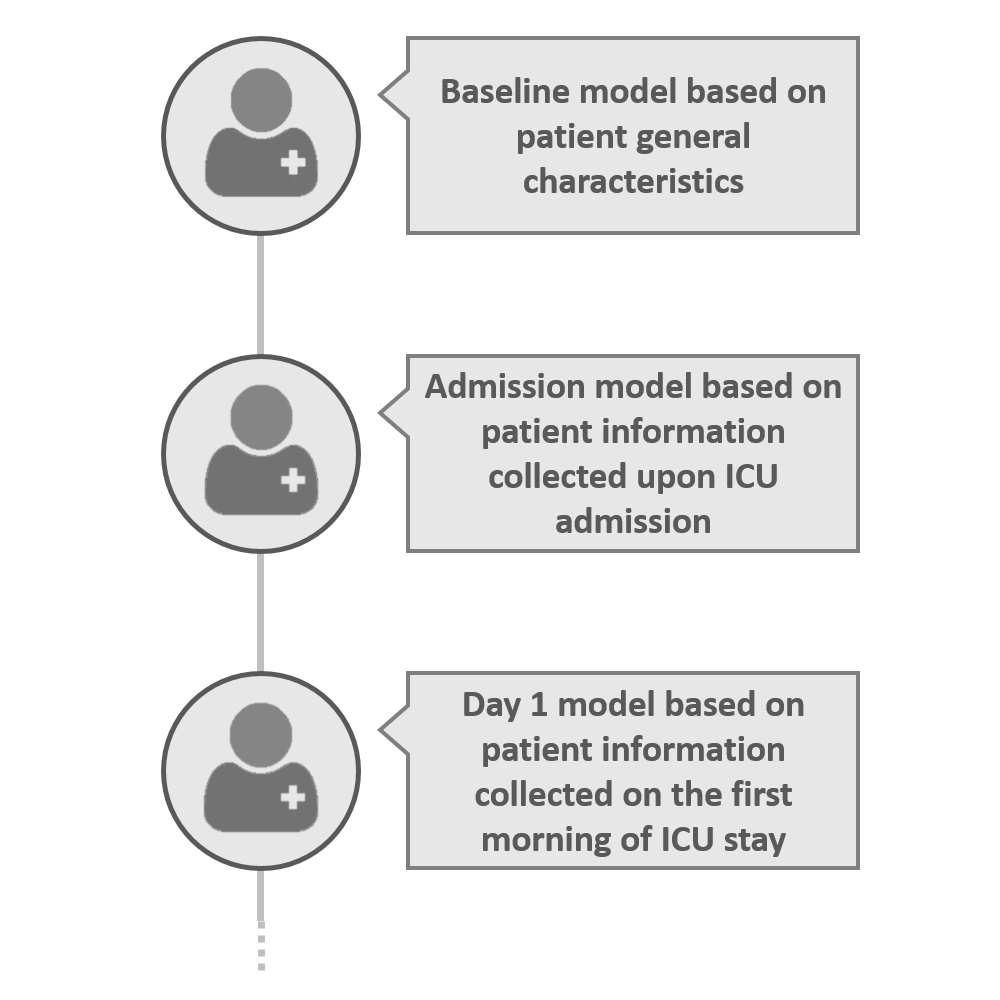
Three models have been developed according to the timing of availability of clinical information of a critically ill patient in an ICU: a baseline model using only demographic data and data known before ICU admission; an admission model that adds data available upon ICU admission; and a day 1 model that, in addition, uses data available on the first day in ICU. This way, the risk of developing AKI can be estimated before ICU admission, at ICU admission, and after the first 1-24 hours in the ICU.
The parameters used by the models are believed to be available in most ICU patients throughout the world. In addition the models are sparse and use a relatively low amount of attributes. Nevertheless, because they been developed and validated in the EPaNIC database, the performance and validity of the model in datasets from other centers might be different from the performance in the original EPaNIC dataset. The presented models are a work in progress, and can be further validated and recalibrated using more multicenter data. The models requires prospective external validation.
1. Casaer Michael P., et al. "Early versus late parenteral nutrition in critically ill adults." N Engl J Med 365.6 (2011): 506-517.
Our research group
Our group combines a research laboratory with a large intensive care unit (65 beds, over 2500 patients each year), both located on the Health Sciences Campus of Gasthuisberg in Leuven, Belgium. This unique setting allows a very fast and effective interaction among clinicians and basic researchers within the research team, allowing research from “bed to bench and from bench to bed”.